New RTG1721 Publication: Chromatin Structure
Slip-sliding away …
04.09.2018
The DNA in the cell nucleus is highly condensed, and genes must be rendered accessible before they can be activated. Karl-Peter Hopfner (RTG1721), Kilian Knoll (RTG1721), Philipp Korber and coworkers have now described the action of a protein complex that serves as a yardstick to measure lengths of exposed DNA.
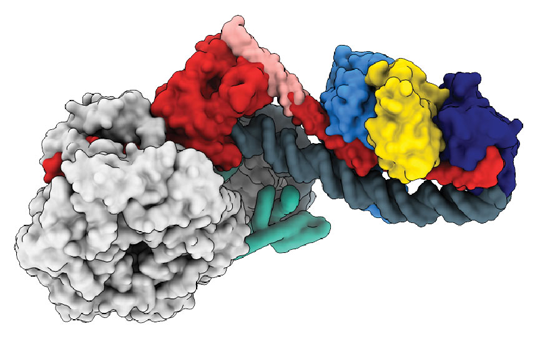
Structural Model of the INO80-Remodeller (Hopfner-Lab/LMU, Nature)
In the cell nucleus, the genomic DNA is packaged into a tightly condensed form, which is referred to as chromatin. The basic unit of chromatin organization is the nucleosome, a DNA-protein complex consisting of a defined length of DNA wrapped around a bead-like structure which is made of histone proteins. The individual nucleosomes are connected by a short length of linker DNA, forming a string of beads which are in turn packed together by special cross-linking proteins. The detailed structure of chromatin regulates access to the genes, and therefore plays a vital role in the control of gene expression. In response to metabolic signals, environmental changes and developmental processes, mechanisms are triggered that dynamically modify chromatin structure, making genes accessible for activation whose products enable the cell to adapt to changes in local conditions. Large protein complexes called chromatin remodellers play a central role in this process. These molecular machines act locally to disrupt the contacts between nucleosomal histones and DNA and effectively slide the nucleosome along the DNA. Karl-Peter Hopfner's lab in collaboration with a team around Dr. Philipp Korber at the Biomedical Center, has dissected the function of an important module of the remodeller INO80 in this process. The new findings have just been published in the journal Nature Structural & Molecular Biology.
More information please visit LMU.de/news
Original Publication:
The nuclear actin-containing Arp8 module is a linker DNA sensor driving INO80 chromatin remodeling.
Knoll KR, Eustermann S, Niebauer V, Oberbeckmann E, Stoehr G, Schall K, Tosi A, Schwarz M, Buchfellner A, Korber P & Hopfner KP
Nature Structural & Molecular Biology. 2018 Sept 3. doi: 10.1038/s41594-018-0115-8.
See also:
Structural Biology: The art of unpacking

